3.2. Semantic interoperability1
3.2.1. The problem of semantic interoperability
Semantic interoperability goes beyond shared data formats or common terminologies — it requires that different systems interpret exchanged data in the same way. Achieving this depends on how well the ontology of each system captures the intended meaning of the concepts it uses. The following diagrams illustrate how systems can differ in their approach to a shared conceptualisation, and how these differences affect their ability to collaborate meaningfully.
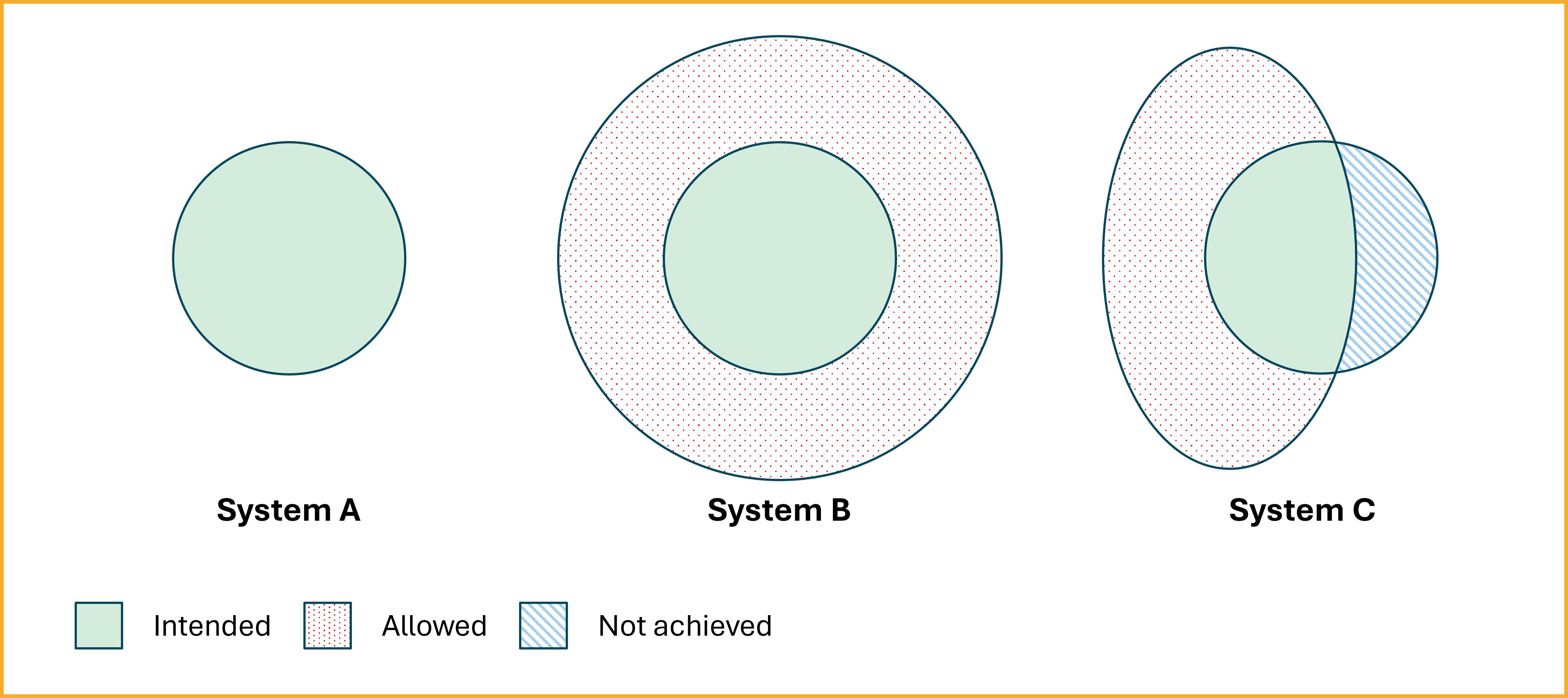
The diagram above illustrates how three different systems attempt to represent the same conceptualisation — i.e. a shared understanding of the real world — but with different levels of success. Each system uses a different ontology to constrain its internal model, leading to differences in how faithfully they capture the intended meaning.
-
System A (left) represents an ideal case. Its ontology closely aligns with the intended semantics (green area), without adding unintended interpretations or excluding valid ones. This is described as a close approximation of the intended models — a highly desirable, though often difficult to achieve, situation for semantic interoperability.
-
System B (middle) uses a more permissive ontology. The red dotted area shows unintended models that are logically consistent with System B's formalism but deviate from the intended meaning. Such over-generalisation can lead to false agreement: systems that appear interoperable because they use the same vocabulary but actually interpret the terms differently.
-
System C (right) makes the opposite mistake. Its ontology is too restrictive: the blue striped area represents valid meanings it does not account for. This can happen when a system's constraints are too narrow or incomplete, leading to the exclusion of necessary interpretations and loss of information.
True semantic interoperability therefore requires more than syntactic or logical alignment. It requires a shared worldview — a convergence on what exists, how it can be described, and which interpretations are valid.
3.2.2. How is semantic interoperability achieved?
Achieving semantic interoperability involves several steps:
- Standardised vocabularies and ontologies: Using shared terminologies such as SNOMED CT, LOINC, or domain-specific ontologies to ensure consistent understanding of data elements.
- Metadata and annotations: Adding semantic layers to data using RDF, OWL or JSON-LD to provide context and meaning.
- Mappings and alignments: Creating links between different vocabularies or datasets to facilitate data integration and interoperability.
- Tools and platforms: Using interoperability frameworks, APIs and knowledge graphs to support seamless data exchange and understanding.
- Ontologically grounded languages: Using modelling languages that help to make the ontological assumptions embedded in data and systems explicit. As Guizzardi argues, it is not enough that representation languages are formally expressive — they must support users in articulating and verifying the actual semantics of their models through ontological commitments. OntoUML is one such language — explicitly grounded in the Unified Foundational Ontology (UFO) — that enables semantically rich, internally consistent and verifiable conceptual modelling.
3.2.3. Implications for data stations
Generic vs. specific: a spectrum of mapping effort
Not all semantic interoperability problems are equally hard. At one end of the spectrum sits generic, broadly shared clinical data — patient demographics, diagnoses, medications, encounters, procedures — for which well-established standards (FHIR, OMOP CDM, SNOMED CT, LOINC, RxNorm) and increasingly mature cross-model mappings already exist. At the other end sit highly specific research domains — radiomics, multi-omics, specialised biobanking — where the conceptual models are still contested, coding systems are sparse, and achieving semantic traceability requires substantial domain-expert effort. Data station operators must locate themselves on this spectrum and choose an implementation strategy accordingly. For generic data, the investment in reusing or extending existing mappings pays off quickly. For domain-specific research questions, the effort of detailed ontological mapping is unavoidable and, until sufficient maturity is reached, routing data through a central SPE for expert integration is often the more pragmatic choice 2.
Existing mappings for common clinical data
For common clinical data, a growing ecosystem of mappings already exists in the Netherlands and internationally. SSSOM (Simple Standard for Sharing Ontology Mappings) combined with SEMAPV (Semantic Mapping Vocabulary) provides a standardised, machine-readable format for expressing these mappings, including their provenance, confidence, and justification. Existing mappings could be expressed and maintained using these standards, for example:
- DHD Diagnosis and Procedures Thesaurus — a Dutch hospital coding thesaurus with potential for standardised alignment to ICD-10 and SNOMED CT.
- Z-Index to RxNorm — ErasmusMC has produced a mapping from the Dutch Z-Index medication register to RxNorm; Cornet et al. describe the full chain from Z-Index → ATC → RxNorm.
- FHIR R4 ↔ OMOP CDM 5.4 — the most mature cross-model mapping in the international clinical data ecosystem, formalised in the HL7 FHIR-OMOP Implementation Guide.
In scaling up data stations, we imagine a national collaboration to express existing Dutch mappings in SSSOM + SEMAPV to make them reusable, citeable and auditable across participating institutions.
A Rosetta Stone for healthcare data models
Taking inspiration from the Rosetta Stone, we envisage an artefact that made it possible to create (semi-)automatic translation between different healthcare information models in anpreviously incompatible writing systems. As an example, plugin-rosetta propopses an implementation of a generic translation layer between FHIR R4 and OMOP CDM 5.4. It illustrates concretely how a Rosetta Stone for generic clinical data can be built and maintained.
The implementation rests on three pillars:
-
LinkML schemas as the common schema layer. Both the FHIR R4 resources and the OMOP CDM 5.4 tables are described as LinkML schemas. LinkML provides a technology-neutral, human-readable schema language that can be compiled to JSON Schema, RDF, OWL, Pydantic models, and more. Expressing both models in the same schema language makes the mapping surface explicit and auditable.
-
Annotation of mappings as machine-readable mapping records. Within the LinkML schema for FHIR resources, each mapped field carries an mappping annotation that clarifies whether the mapping to its OMOP counterpart is exact, narrow or broad. These annotations correspond directly to SSSOM's
skos:typeMatchpredicate, grounding the mapping in a recognised standard vocabulary:# fhir_resources.yaml (excerpt) slots: gender: exact_mappings: - omop:gender_concept_id - omop:gender_source_valueThe mappings are derived from the standard HL7 FHIR-OMOP IG FML structure maps such that every mapping decision traces back to an authoritative, versioned upstream source rather than an ad-hoc field-by-field judgement.
-
Three-layer validation. Output is validated at the columnar level (schema constraints), the record level (Pydantic v2 models generated from LinkML), and the semantic level (LinkML
validate()). This layered approach catches structural errors, type mismatches, and semantic constraint violations separately, making it easier to diagnose and fix integration issues.
The same Rosetta Stone pattern — LinkML schemas for both source and target models, annotation of mappings grounded in SSSOM predicates, authoritative upstream source code as the source of truth — can be applied to other common model pairs in the Dutch health data landscape, such as ZIB ↔ OMOP or DHD ↔ SNOMED CT.
Deep ontological alignment for specific research questions
Where the Rosetta Stone approach gives a pragmatic, reusable solution for generic data, specific research questions with complex or contested semantics require a deeper level of ontological work. The Health-RI Semantic Interoperability Initiative provides a rigorous framework for this harder problem.
The framework is built around the Health-RI Ontology (HRIO), developed using a model-driven architecture that separates meaning from representation:
- HRIO OntoUML (CIM): a conceptual model at the computation-independent layer, grounded in the Unified Foundational Ontology (UFO). Domain experts use this layer to validate that the intended meaning of concepts is captured correctly, before any computational commitments are made.
- HRIO gUFO/OWL (PIM): a machine-readable OWL 2 DL implementation of the same meanings at the platform-independent layer, suitable for reasoning and Semantic Web integration. Every OWL class inherits its semantics from its OntoUML counterpart, ensuring that meaning and representation remain traceable across layers.
External ontologies, terminologies, and schemas are aligned to HRIO through the Health-RI Mapping Vocabulary (HRIV), which provides a meaning-level mapping layer on top of SSSOM and SKOS:
hriv:hasExactMeaning— the external concept's intended semantics are fully and precisely defined by the linked HRIO concept (exactly one allowed per external concept; entailsskos:exactMatch).hriv:hasBroaderMeaningThan— the external concept is broader in scope than the linked HRIO meaning (entailsskos:narrowMatch).hriv:hasNarrowerMeaningThan— the external concept is narrower in scope than the linked HRIO meaning (entailsskos:broadMatch).
This meaning-level vocabulary avoids the "false agreement" trap described in section 3.2.1: two standards may share a label like "Man" while embedding different conceptualisations (karyotype-based vs. gender-based). HRIV mappings make the intended semantic commitment explicit, rather than relying on label similarity alone.
The architectural payoff is a semantic hub: instead of maintaining N×M pairwise mappings between all combinations of standards and local schemas, each external artifact maps once to HRIO. When two external terms map to the same HRIO meaning, their intended semantics become comparable through the shared reference — without requiring a bespoke pairwise mapping for every combination. All mapping artifacts are published with stable, citable persistent identifiers (w3id), versioned using append-only supersession (never overwritten), and governed through a role-separated review process (Mapper ≠ Reviewer; Curator publishes).
For specific domains such as radiomics, achieving this level of semantic interoperability requires very specialised knowledge 3. Until sufficient ontological maturity is reached in such domains, routing data through a central SPE for expert integration remains the more pragmatic option.
-
The text on this page is largely derived from the Semantic Interoperability initiative of Health-RI. ↩
-
Varsha Gouthamchand, Ananya Choudhury, Frank J P Hoebers, Frederik W R Wesseling, Mattea Welch, Sejin Kim, Joanna Kazmierska, Andre Dekker, Benjamin Haibe-Kains, Johan van Soest, and Leonard Wee. Making head and neck cancer clinical data Findable-Accessible-Interoperable-Reusable to support multi-institutional collaboration and federated learning. BJR\textbar Artificial Intelligence, 1(1):ubae005, March 2024. URL: https://doi.org/10.1093/bjrai/ubae005 (visited on 2025-09-15), doi:10.1093/bjrai/ubae005. ↩
-
Petros Kalendralis, Zhenwei Shi, Alberto Traverso, Ananya Choudhury, Matthijs Sloep, Ivan Zhovannik, Martijn P.A. Starmans, Detlef Grittner, Peter Feltens, Rene Monshouwer, Stefan Klein, Rianne Fijten, Hugo Aerts, Andre Dekker, Johan van Soest, and Leonard Wee. FAIR-compliant clinical, radiomics and DICOM metadata of RIDER, interobserver, Lung1 and head-Neck1 TCIA collections. Medical Physics, 47(11):5931–5940, 2020. _eprint: https://aapm.onlinelibrary.wiley.com/doi/pdf/10.1002/mp.14322. URL: https://onlinelibrary.wiley.com/doi/abs/10.1002/mp.14322 (visited on 2026-01-09), doi:10.1002/mp.14322. ↩